I will start the blog with a statement: "Mitochondrial Disorders are not only due to mutations in the mitochondrial genome but also due to mutations in the nuclear genome."
While I justify my statement, I will explain the Mitochondrial Genomics. Mitochondria are the powerhouses of the cell. They break down components from cellular metabolism into ATP (Chemical Energy) using oxygen.
Fun Fact - Some believe mitochondria are bacteria that eventually enter the cells and undergo a symbiotic relationship with the nuclear genome, sharing some of its genes with the human nuclear genome. The circular nature of mitochondrial DNA (mtDNA) supports this theory.
Besides, 40% of the proteins coded by mitochondria are bacterial. Our CCMB team, led by Dr.Shankaranarayana, supports this theory with solid evidence.
Mitochondrial Disorders:
Mitochondrial disorders are complex disorders that arise due to dysfunction of the mitochondrial respiratory chain, which is the final pathway for aerobic metabolism (generating energy in the presence of oxygen). So, the mitochondrial disorder affects any tissue or organ ( the eye, ear, liver, renal tissues, etc.) that is highly dependent on aerobic mechanisms.
Common clinical features of mitochondrial disorders include:
- Ptosis
- External ophthalmoplegia
- Proximal myopathy
- Exercise intolerance
- Cardiomyopathy
- Sensorineural deafness
- Optic atrophy
- Pigmentary retinopathy
- Diabetes mellitus
Primary and Secondary Mitochondrial Disorders
The following are the mitochondrial disorders:
Primary Mitochondrial Disorders (PMD)– Pathogenic variants in the mitochondrial respiratory chain and related peptides cause primary mitochondrial disorders (PMD). MitochondrialDNA is 16kb in length with 37 genes. Mutations in these genes disturb the function of mitochondria leading to disease. Approximately 1500 nuclear genes are involved in coding respiratory chain subunits and proteins for mtDNA maintenance. Mutations in these 1500 nuclear genes can also cause mitochondrial disorders.
Secondary Mitochondrial Disorders (SMD)–Disorders that affect mitochondrial mechanisms and their functions cause secondary mitochondrial disorders (SMD). The inheritance of mitochondria is also unique. SMD is primarily maternal because the sperm's mitochondrial DNA is degraded in the urinary tract (except for some rare cases).

Importance of MtWGS-NGS Sequencing
Each cell contains thousands of mitochondria. The severity of the mitochondrial disorder depends on the number of defective mitochondrial DNA present in each cell. Besides, the mitochondrial disorder is tissue-specific, meaning we need to select the exact tissue to diagnose the mutations. Many mutations present in the affected individuals' muscle cells are not shown in blood cells, thus making the diagnosis of mitochondrial disorders more complex.
The complexity in mitochondrial genetics is due to complicated symptoms, inheritance patterns, expression patterns, DNA copies, and types of genomic changes. Moreover, mitochondrial genetics is complicated due to mitochondria’s involvement in energy generation. The traditional way of identifying mitochondrial disorders is to understand the symptoms and screen for hotspot mutations and deletions, followed by Sanger sequencing. However, the involvement of many genes and their length makes the diagnosis difficult to establish.
Mitochondrial sequencing with Next Generation Sequencing (NGS) can identify the structural variants and mutations in a single step. Clinicians are more interested in doing Whole Exome Sequencing (WES) with Mitochondrial DNA Whole Genome Sequencing (WES+Mitochondria-WGS) because most mitochondrial disorders are due to the mutations in the nuclear genome. It is essential to recognize that mitochondrial genetics involves two separate genomes, mitochondrial and nuclear. Thus a genetic evaluation of the patient with maternal and mendelian inheritance with a thorough one-step diagnosis is necessary.
Mapmygenome’s Next Generation Sequencing Solutions
Mapmygenome Next Generation Sequencing (NGS) Solutions give extensive coverage of all genes. It comes with high precision analysis, detailed interpretation, and easy-to-read clinical reports. These NGS solutions help physicians to make accurate decisions for diagnosis, treatment, and management of several health conditions. Mapmygenome’s NGS solutions include Whole Exome Sequencing(WES) and Whole Genome Sequencing (WGS)
Advantages of MMG’s WES and WGS
Mitochondrial Panel in MMG’s WES
Mapmygenome has designed a panel in Whole Exome Sequencing (WES) to evaluate the mitochondrial and nuclear DNA. This panel analyses the entire exome(~23,000 genes) and the 37 genes on the mitochondrial DNA(16kb) with 100-120X coverage to identify the structural variants and mutations involved in Mitochondrial disorders.
Whole Exome Sequencing
Mapmygenome’s Whole Exome Sequencing (MMG WES) analyses the entire exome(~23,000 genes) with 100-120X coverage. It is ideal for the diagnosis of multi-gene disorders. MMG’s WES also includes comprehensive bioinformatics analysis, clinical analysis, and detailed report.
MMG’s WES helps in:
Prognosis: MMG’s WES gives healthy individuals improved risk assessment and preventive strategies for common diseases such as diabetes, heart diseases, and cancer.
Molecular Diagnosis: MMG WES helps in confirming a clinically diagnosed genetic condition. The report provides a genetic diagnosis that can inform appropriate treatment, management, inheritance, accurate recurrence risks, and reproductive decision-making.
Discovery: In cases of no diagnosis but clinical presentation suggestive of a genetic condition, MMG’s WES can help confirm it.
Efficiency: Cost-effective and time-effective solution (compared against sequential single-gene testing) for cases where a change in one of the genes may result in a specific clinical picture.
Utility: All genes covered. Tests for single-gene and multi-gene conditions. Ideal for targeted management of complex syndromes, epilepsy, ASDs, neuropathy, and heterogeneous phenotypes.
Whole Genome Sequencing
Mapmygenome’s Whole Genome Sequencing(MMG’s WGS) analyses 98% of the human DNA. It means WGS identifies the variants in the entire genome rather than a few selected regions with 30X coverage. WGS is useful in Diagnostics, Wellness, Fitness, Nutrition, and Medication. It also includes comprehensive bioinformatics analysis, clinical analysis, and a detailed report.
MMG’s WGS helps in:
Making a diagnosis: Some genetic conditions can pose a diagnostic challenge. A single gene testing cannot provide an in-depth analysis of complex diseases. MMG’s WGS can identify complex and rare conditions and help in their management.
Precision Medicine: MMG’s WGS analyses changes in DNA and help in identifying medication that is effective for a person. It significantly changes the approach to treatment for diseases such as cancer, diabetes, heart diseases etc.
Disease Prevention: Genetic predispositions for diseases are encoded in DNA. MMG’s WGS can identify such predispositions to implement a strategic prevention plan.
The process in Mapmygenome’s NGS solutions includes:
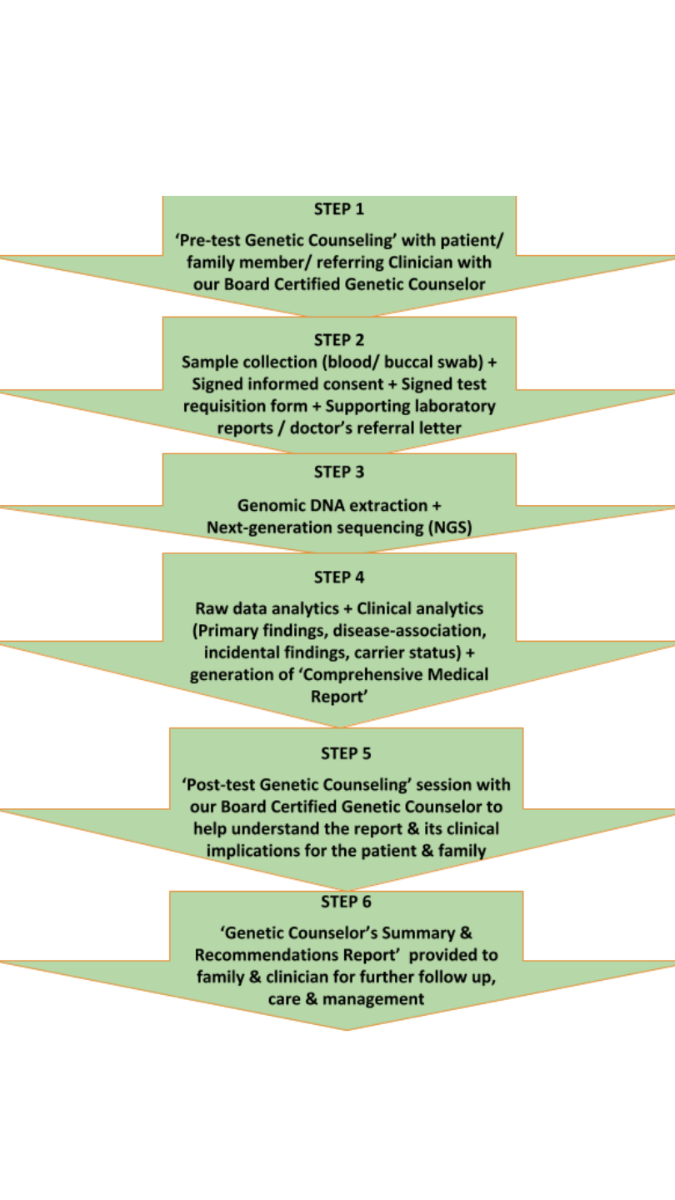
1. Pre-test genetic counselling with patients, family members, and clinicians with Mapmygenome’s Board Certified Genetic Counselor.
2. Sample Collection( blood/ buccal swab). The patient’s doctor places a request for the test. The sample is tested for susceptibility.
3. Using Next Generation Sequencing(NGS), genetic data is subjected to extensive analysis to extract health information and generate a comprehensive health report.
4. Post-test genetic counseling session with our genetic counselor to help understand the report and its clinical implications for the patient family
5. Clinicians and families receive a genetic counselor’s summary and recommendations report for further follow-up and management of health conditions.
Click here to know more about MMG's Whole Exome/Genome Sequencing Solutions

